Research Symposium
26th annual Undergraduate Research Symposium, April 1, 2026
Martina Madrzak Poster Session 4: 3:00 pm - 4:00 pm / Poster #2
BIO
Martina Madrzak is a sophomore at Florida State University pursuing a Bachelor of Science in Biological Sciences. Martina was first inspired by scientific research when she attended a Duke University Summer Research Program following her freshman year, where she studied Cryptococcus neoformans, a fungal pathogen, examining its virulence traits in natural environments and drawing connections to clinical cases. Her work contributed to identifying a potential genetic mutation that enables diploid Cryptococcus cells to sustain infection in the body. Following her experience at Duke, Martina conducted research through FSU's Undergraduate Research Opportunity Program (UROP) with the Solís Lab, focusing on structural biology and biochemistry with a focus on the alpha-actinin-2 protein.
Throughout all this research, Martina has found her enjoying dry lab aspects and has a growing curiosity towards computational biology. Martina will be pursuing an NSF-funded Research Experience for Undergraduates (REU) through Rosetta Commons in the Strauch Lab, where she is working on computational protein engineering and expanding her expertise in structural biology and protein research from the Solís Lab. She is also a member of the Modeling and Design Team of FSU's iGEM team and serves as an officer of the Business and STEM Association.
Martina is passionate about the intersection of computational biology, business, and entrepreneurship, and hopes to pursue opportunities that merge scientific innovation with real-world application.
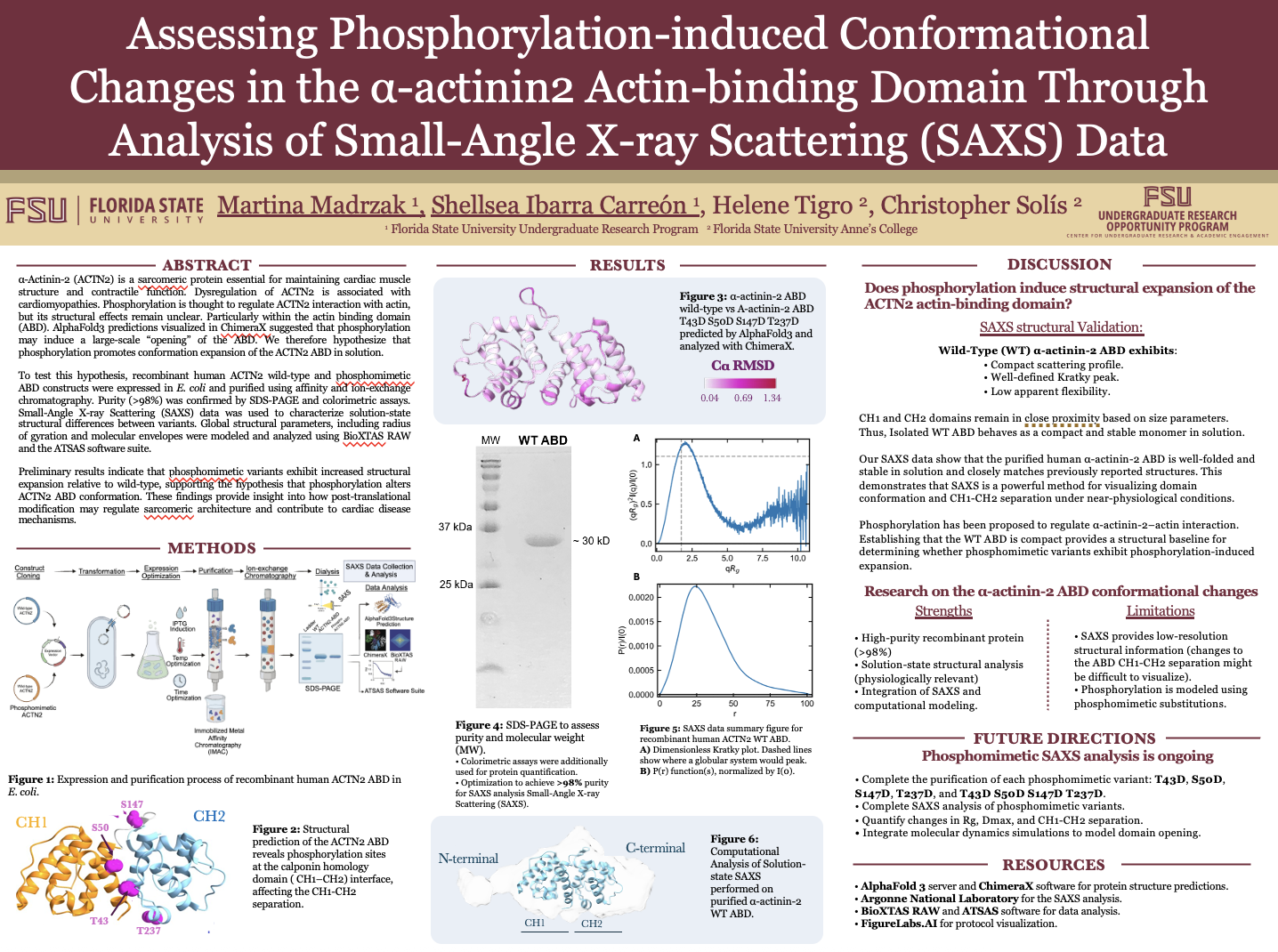
Assessing phosphorylation-induced conformational changes in the α-actinin2 actin-binding domain through analysis of Small-Angle X-ray Scattering (SAXS) data
Authors: Martina Madrzak, Helene TigroStudent Major: Biological Sciences
Mentor: Helene Tigro
Mentor's Department: Department of Health, Nutrition, and Food Sciences Mentor's College: Anne's College Co-Presenters: Shellsea Ibarra Carreón
Abstract
α-Actinin-2 (ACTN2) is a sarcomeric actin-crosslinking protein essential for maintaining cardiac
muscle structure and function, and mutations or dysregulation of ACTN2 have been linked to
cardiomyopathies. Phosphorylation is thought to regulate ACTN2 activity, but its structural
effects remain unclear. Structural predictions generated using AlphaFold3 and visualized in
ChimeraX suggested that phosphorylation may induce an opening of the actin-binding domain
(ABD). Based on these computational observations, we hypothesized that phosphorylation
promotes conformational expansion of the ACTN2 ABD in solution.
To test this, recombinant human ACTN2 wild-type and phosphomimetic ABD constructs were
expressed in Escherichia coli and purified using sonication, ultracentrifugation, affinity chromatography, ion-exchange chromatography, and dialysis. Protein purity (>98%) was verified
by SDS-PAGE and colorimetric protein quantification assays. Small-Angle X-ray Scattering
(SAXS) was then used to examine solution-state structural differences between variants. SAXS
data were processed and analyzed using BioXTAS RAW and the ATSAS software suite to
evaluate global structural parameters, including radius of gyration and molecular envelopes.
Preliminary results indicate that phosphomimetic variants exhibit increased structural expansion
relative to wild-type, supporting the hypothesis that phosphorylation alters ACTN2 conformation.
These findings provide insight into how post-translational modification may regulate sarcomeric
architecture and contribute to cardiac disease mechanisms
Keywords: biology, computational, biochemistry,