Research Symposium
26th annual Undergraduate Research Symposium, April 1, 2026
Gilberto Torres-Reyes Poster Session 2: 10:45 am - 11:45 am / Poster #226

BIO
I am an undergraduate student from Miami, Florida with strong interests in biomedical research and physician–scientist training. Through my studies and research experiences, I am learning to use computational and molecular approaches to better understand the biological mechanisms that drive disease. I am also completing a Directed Individual Study in the Department of Biological Science under the mentorship of Dr. Qian Yin, where I work on the visualization and analysis of cross-linking mass spectrometry data. I plan to pursue an MD/PhD to integrate clinical medicine with biomedical research, with the ultimate goal of contributing to innovative projects that translate scientific discovery into advances that improve health at the population level.
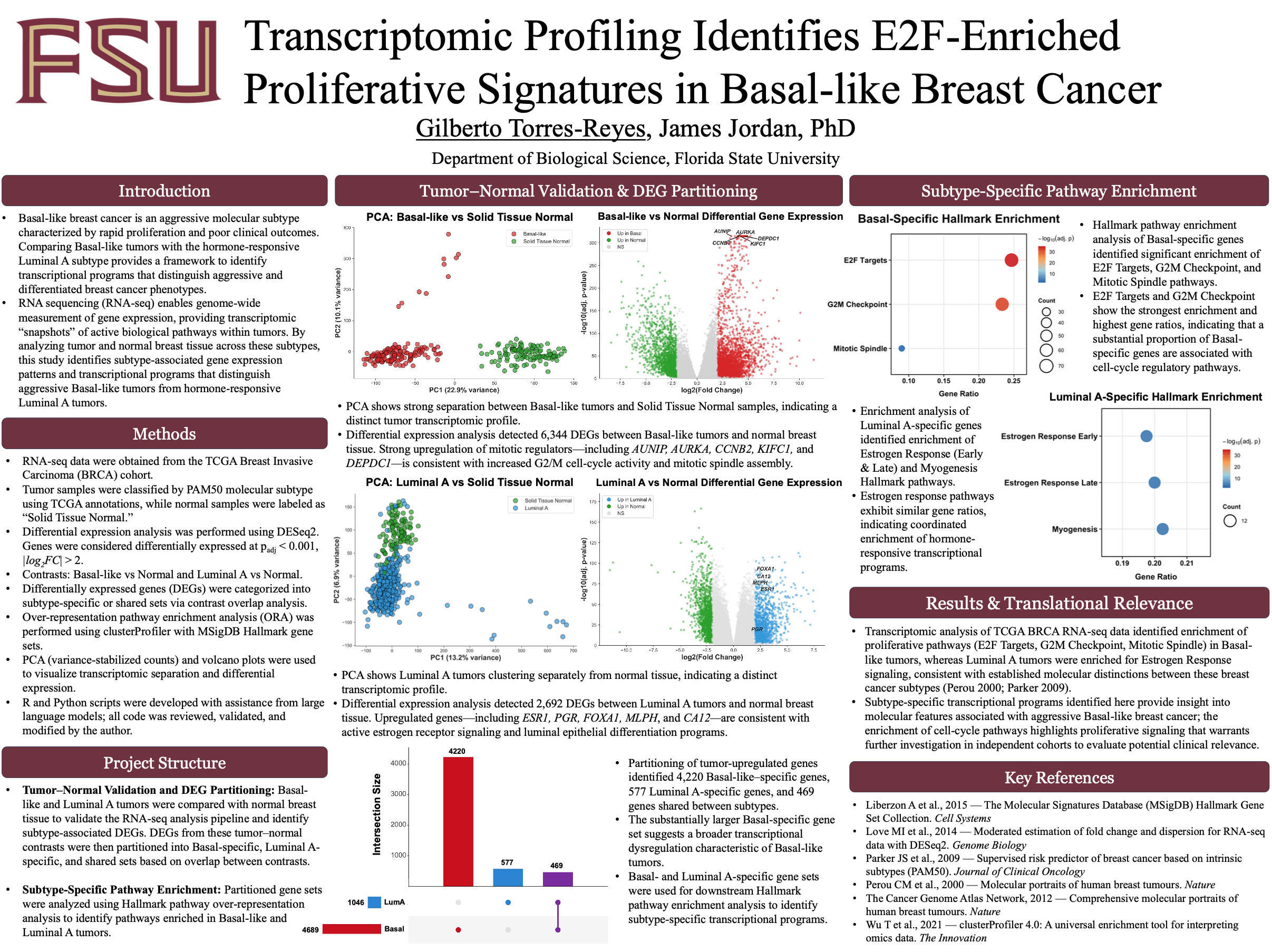
Transcriptomic Profiling Identifies E2F-Enriched Proliferative Signatures in Basal-like Breast Cancer
Authors: Gilberto Torres-Reyes, James Jordan, PhDStudent Major: Public Health, Minor in Chemistry
Mentor: James Jordan, PhD
Mentor's Department: Department of Biological Science Mentor's College: College of Arts and Sciences Co-Presenters:
Abstract
Basal-like breast cancer is an aggressive molecular subtype characterized by rapid proliferation and poor clinical outcomes. To better understand the transcriptional programs associated with this phenotype, RNA sequencing (RNA-seq) data from the TCGA Breast Invasive Carcinoma (BRCA) cohort were analyzed to compare Basal-like and Luminal A breast cancer subtypes with normal breast tissue. Differential gene expression analysis was performed using DESeq2, and transcriptomic patterns were visualized using principal component analysis and volcano plots. Differentially expressed genes were partitioned into Basal-like–specific, Luminal A–specific, and shared gene sets, followed by Hallmark pathway enrichment analysis using clusterProfiler. This analysis identified 4,220 Basal-like–specific genes and 577 Luminal A–specific genes, indicating broader transcriptional dysregulation in Basal-like tumors. Enrichment analysis revealed strong activation of proliferative pathways—including E2F Targets, G2M Checkpoint, and Mitotic Spindle—in Basal-like tumors, whereas Luminal A tumors showed enrichment of estrogen response signaling pathways. These findings highlight distinct transcriptional programs underlying aggressive Basal-like breast cancer and provide insight into biological pathways associated with subtype-specific tumor behavior.
Keywords: Breast cancer, Basal-like breast cancer, RNA-seq analysis, differential gene expression, pathway enrichment analysis, transcriptomics