Research Symposium
26th annual Undergraduate Research Symposium, April 1, 2026
Keyse Kelly Ferraz Poster Session 4: 3:00 pm - 4:00 pm / Poster #188

BIO
Keyse Kelly Ferraz is an undergraduate honors student at Florida State University pursuing a Bachelor of Science in Biological Sciences on the pre-medical track. She is currently in her second year and is involved in undergraduate research through the Undergraduate Research Opportunity Program (UROP). Originally from Brazil, Keyse later moved to Orlando, Florida, where she developed a strong interest in science and medicine.
Her research focuses on analyzing structural changes in cardiac muscle proteins associated with mutations in ACTN2, using computational tools to examine differences in domain organization that may contribute to cardiac disease. She conducts this research under the mentorship of graduate researcher Rhiannon Crawford. In addition to her research, Keyse is actively involved in campus organizations and community engagement initiatives. After completing her undergraduate studies, she plans to attend medical school and pursue a career as a physician while continuing to contribute to research that advances human health.
Analysis of the alpha-actinin-2 actin-binding domain using AI programs
Authors: Keyse Kelly Ferraz, Rhiannon CrawfordStudent Major: Biological Sciences
Mentor: Rhiannon Crawford
Mentor's Department: Health, Nutrition, & Food Sci (HUM_NIP) 135000 Mentor's College: College of Health and Human Sciences Co-Presenters:
Abstract
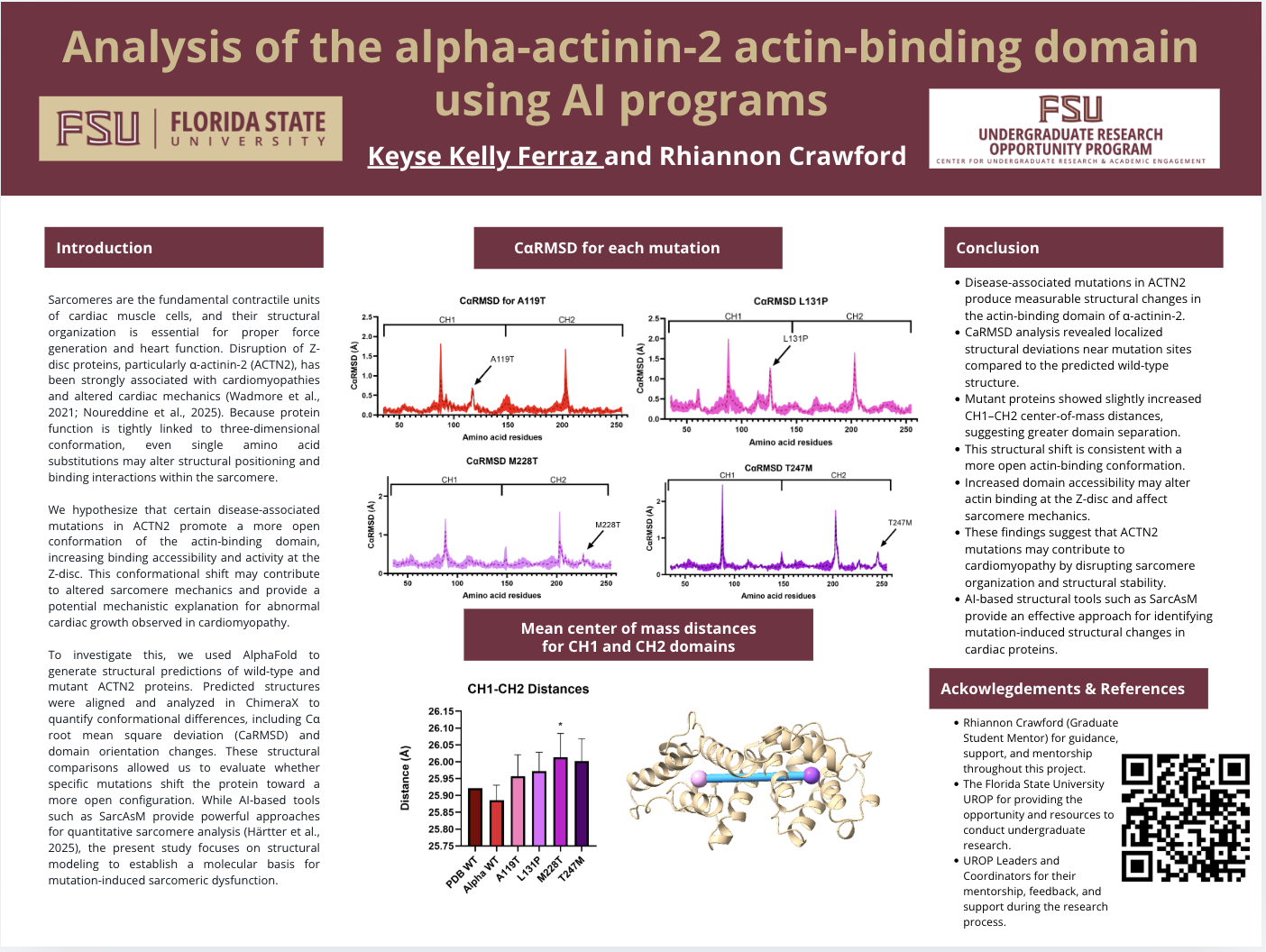
Sarcomeres are the fundamental contractile units of cardiac muscle cells, and their structural organization is essential for proper force generation and heart function. Disruption of Z-disc proteins, particularly α-actinin-2 (ACTN2), has been strongly associated with cardiomyopathies and altered cardiac mechanics (Wadmore et al., 2021; Noureddine et al., 2025). Because protein function is tightly linked to three-dimensional conformation, even single amino acid substitutions may alter structural positioning and binding interactions within the sarcomere.
We hypothesize that certain disease-associated mutations in ACTN2 promote a more open conformation of the actin-binding domain, increasing binding accessibility and activity at the Z-disc. This conformational shift may contribute to altered sarcomere mechanics and provide a potential mechanistic explanation for abnormal cardiac growth observed in cardiomyopathy.
To investigate this, we used AlphaFold to generate structural predictions of wild-type and mutant ACTN2 proteins. Predicted structures were aligned and analyzed in ChimeraX to quantify conformational differences, including Cα root mean square deviation (CaRMSD) and domain orientation changes. These structural comparisons allowed us to evaluate whether specific mutations shift the protein toward a more open configuration. While AI-based tools such as SarcAsM provide powerful approaches for quantitative sarcomere analysis (Härtter et al., 2025), the present study focuses on structural modeling to establish a molecular basis for mutation-induced sarcomeric dysfunction.
Keywords: Computational Biology, Cardiac Muscle, Mutation